Principal component analysis (PCA) is routinely employed on a wide range of problems. From the detection of outliers to predictive modeling, PCA has the ability of projecting the observations described by  variables into few orthogonal components defined at where the data ‘stretch’ the most, rendering a simplified overview. PCA is particularly powerful in dealing with multicollinearity and variables that outnumber the samples (
variables into few orthogonal components defined at where the data ‘stretch’ the most, rendering a simplified overview. PCA is particularly powerful in dealing with multicollinearity and variables that outnumber the samples (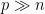 ).
).
It is an unsupervised method, meaning it will always look into the greatest sources of variation regardless of the data structure. Its counterpart, the partial least squares (PLS), is a supervised method and will perform the same sort of covariance decomposition, albeit building a user-defined number of components (frequently designated as latent variables) that minimize the SSE from predicting a specified outcome with an ordinary least squares (OLS).
Although there is a plethora of PCA methods available for R, I will only introduce two,
- prcomp, a default function from the R base package
- pcaMethods, a bioconductor package that I frequently use for my own PCAs
 REGISTER FOR FREE WEBINAR
X
REGISTER FOR FREE WEBINAR
X
 Thank you for registering
Join Edureka Meetup community for 100+ Free Webinars each month
JOIN MEETUP GROUP
Thank you for registering
Join Edureka Meetup community for 100+ Free Webinars each month
JOIN MEETUP GROUP