Data Science with Python Certification Course
- 131k Enrolled Learners
- Weekend
- Live Class
R for Data Science is a must learn for Data Analysis & Data Science professionals. With its growth in the IT industry, there is a booming demand for skilled Data Scientists who have an understanding of the major concepts in R. One such concept, is the Decision Tree.
In this blog we will discuss :
1. How to create a decision tree for the admission data.
2. Use rattle to plot the tree.
3. Validation of decision tree using the ‘Complexity Parameter’ and cross validated error.
4. Prune the tree on the basis of these parameters to create an optimal decision tree.
To understand what are decision trees and what is the statistical mechanism behind them, you can read this post : How To Create A Perfect Decision Tree
To create a decision tree in R, we need to make use of the functions rpart(), or tree(), party(), etc.
rpart() package is used to create the tree. It allows us to grow the whole tree using all the attributes present in the data.
> library("rpart")
> setwd("D://Data")
> data <- read.csv("Gre_Coll_Adm.csv")
> str(data)
'data.frame': 400 obs. of 5 variables:
$ X : int 1 2 3 4 5 6 7 8 9 10 ...
$ Admission_YN : int 0 1 1 1 0 1 1 0 1 0 ...
$ Grad_Rec_Exam: int 380 660 800 640 520 760 560 400 540 700 ...
$ Grad_Per : num 3.61 3.67 4 3.19 2.93 3 2.98 3.08 3.39 3.92 ...
$ Rank_of_col : int 3 3 1 4 4 2 1 2 3 2 ...
> View(data)> adm_data<-as.data.frame(data) > tree <- rpart(Admission_YN ~ adm_data$Grad_Rec_Exam + adm_data$Grad_Per+ adm_data$Rank_of_col, + data=adm_data, + method="class")
rpart syntax takes ‘dependent attribute’ and the rest of the attributes are independent in the analysis.
Admission_YN : Dependent Attribute. As admission depends on the factors score, rank of college, etc.
Grad_Rec_Exam, Grad_Per, and Rank_of_col : Independent Attributes
rpart() returns a Decison tree created for the data.
If you plot this tree, you can see that it is not visible, due to the limitations of the plot window in the R console.
> plot(tree) > text(tree, pretty=0)
To enhance it, let us take some help from rattle :
> library(rattle) > rattle()
Rattle() is one unique feature of R which is specifically built for data mining in R. It provides its own GUI apart from the R Console which makes it easier to analyze data. It has built-in graphics, which provides us better visualizations as well. Here we will use just the plotting capabilities of Rattle to achieve a decent decision tree plot.
> library(rpart.plot) > library(RColorBrewer)
rpart.plot() and RcolorBrewer() functions help us to create a beautiful plot. ‘rpart.plot()’ plots rpart models. It extends plot.rpart and text.rpart in the rpart package. RcolorBrewer() provides us with beautiful color palettes and graphics for the plots.
> fancyRpartPlot(tree)
This was a simple and efficient way to create a Decision Tree in R. But are you sure that this is the optimal ‘Decision Tree’ for this data? If not, the following validation checks will help you.
Meanwhile, if you wish to learn R programming, check out our specially curated course by clicking on the below button.
To validate the model we use the printcp and plotcp functions. ‘CP’ stands for Complexity Parameter of the tree.
Syntax : printcp ( x ) where x is the rpart object.
This function provides the optimal prunings based on the cp value.
We prune the tree to avoid any overfitting of the data. The convention is to have a small tree and the one with least cross validated error given by printcp() function i.e. ‘xerror’.
Cross Validated Error :
To find out how the tree performs, is calculated by the printcp() function, based on which we can go ahead and prune the tree.
> printcp(tree) Classification tree: rpart(formula = Admission_YN ~ adm_data$Grad_Rec_Exam + adm_data$Grad_Per + adm_data$Rank_of_col, data = adm_data, method = "class") Variables actually used in tree construction: [1] adm_data$Grad_Per adm_data$Grad_Rec_Exam adm_data$Rank_of_col Root node error: 127/400 = 0.3175 n= 400 CP nsplit rel error xerror xstd 1 0.062992 0 1.00000 1.00000 0.073308 2 0.023622 2 0.87402 0.92913 0.071818 3 0.015748 4 0.82677 0.99213 0.073152 4 0.010000 8 0.76378 1.02362 0.073760
From the above mentioned list of cp values, we can select the one having the least cross-validated error and use it to prune the tree.
The value of cp should be least, so that the cross-validated error rate is minimum.
To select this, you can make use of this :
fit$cptable[which.min(fit$cptable[,”xerror”]),”CP”]
This function returns the optimal cp value associated with the minimum error.
Let us see what plotcp() function fetches.
> plotcp(tree)
Plotcp() provides a graphical representation to the cross validated error summary. The cp values are plotted against the geometric mean to depict the deviation until the minimum value is reached.
> ptree<- prune(tree, + cp= tree$cptable[which.min(tree$cptable[,"xerror"]),"CP"]) > fancyRpartPlot(ptree, uniform=TRUE, + main="Pruned Classification Tree")
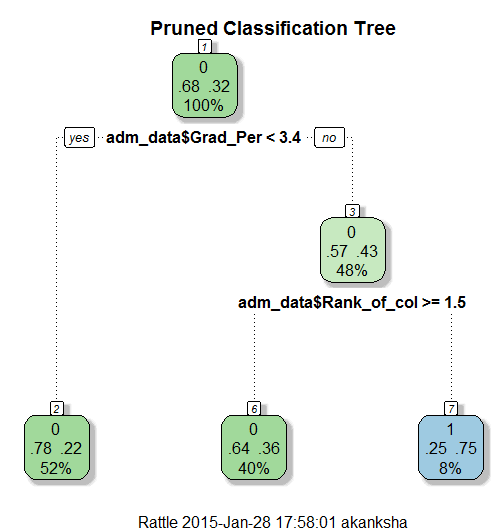
Thus we create a pruned decision tree.
If you wish to get a head-start on R programming, check out the Data Analytics with R course from Edureka.
Got a question for us? Please mention them in the comments section and we will get back to you.
Related Posts:
 Thank you for registering Join Edureka Meetup community for 100+ Free Webinars each month JOIN MEETUP GROUP
Thank you for registering Join Edureka Meetup community for 100+ Free Webinars each month JOIN MEETUP GROUPedureka.co
nice I was just looking how on earth you can prune a tree and I found it
Hi, Thanks for the good article!! I still have a doubt regarding the goodness of the tree.Drawing parallel to the regressions where we check R square value or gain to check the effectiveness of the regression, do we have a similar metric for decision trees. Thanks in advance.
Hi team, can you please share the data to: wasim923@gmail.com
Hey Waism, thanks for checking out our blog!
We have shared it with you. Do subscribe to stay posted on upcoming blogs. Cheers!
hello! where can i donwload the package that you describe here to make this type of tree? thank you
+Joao Marta, thanks for checking out our blog. The packages used in the above example are rpart, rpart.plot,RColorBrewer and rattle. All the these packages can be directly installed through RStudio from the console window through the command:
install.packages(“rpart”)
install.packages(“rpart.plot”)
install.packages(“RColorBrewer”)
install.packages(“rattle”)
Also Try using
install.packages(‘rpart’,dependencies=TRUE) to automatically install all the relevant packages needed for decision tree making process.
Hope this helps. Cheers!
Hi, anyone knows where to download or get the read.csv(“Gre_Coll_Adm.csv”) ?
I encode the data in excel file .csv
Thanks Ma’am =)
“Gre_Coll_Adm.csv”) can u please send this data set to this anandmtech1985@gmail.com
+OJLiwanag, thanks for checking out our blog. Please share your email address with us and we will send it to you. Cheers!
Please send me the data file at asif.karim.csedu@gmail.com
Hi, we have sent you the file. Cheers!
Hi, thanks a million.
“Gre_Coll_Adm.csv”) can u send this data set to this anandmtech1985@gmail.com
“Gre_Coll_Adm.csv”) can u send this data set to this mail id anandmtech1985@gmail.com
please send me the file (“Gre_Coll_Adm.csv”) to ravi62in@yahoo.com;
Hey Ramanathan, thanks for checking out our blog!
We have shared it with you. Do subscribe to stay posted on upcoming blogs. Cheers!
Hi Team,
Can you please send me this file @ abhinavsri23@gmail.com
Hey Abhinav, thanks for checking out our blog!
We have shared it with you. Do subscribe to stay posted on upcoming blogs. Cheers!
Please send the data to mourao2@gmail.com?
Tks much!
Hey Carol, thanks for checking out our blog!
We have shared it with you. Do subscribe to stay posted on upcoming blogs. Cheers!
hi
Can you lease share the dataset (“Gre_Coll_Adm.csv”) ? at ravi62in@yahoo.com
Nice blog
great job !!!
Thanks, Gopesh! You may like this blog too: https://www.edureka.co/blog/implementing-kmeans-clustering-on-the-crime-dataset/. Cheers!
nice blog
great blog!
Thanks, Divya! We’re glad you found the blog useful. Do subscribe to our blog to stay posted on upcoming R programming blogs.
how to easily calculate the accuracy of prediction of test dataset